Krasileva Lab
Full description of lab culture - see Krasileva Lab Charter
231 Koshland Hall, Berkeley CA 94720 kseniak [at] berkeley.edu.
Lab News
📣 We are seeking postdocs interested in comparative immunology with a focus on innate immune systems of plants, fungi + computational biology. We’re excited to host postdoc fellowship applicants in our comparative immunology team. Reach out by email kseniak@berkeley.edu
March 2026
New lab publication out in New Phytologist! The annotated blueprint: integrated functional genomic resources for a model tetraploid wheat cv. Kronos 🌾 chromosome-scale genome, high-conf annotations, manually curated 1000+ NLRs, ~3000 remapped datasets 🔗 doi.org/10.1111/nph....
Past talks (recorded)
Scientific conferences and seminars
Chandler Sutherland - "The Rate, Context, and Consequences of Mutation in the Evolution of the Plant Immune System" - Dissertation Finishing Talk, May 22, 2026 - link to recording (40 min seminar)
Ksenia Krasileva - The Rise of Resistance: Natural Diversity Informs Rational Modifications of Plant Immune Receptors (and also exploration of fungal immune system) - invited seminar at MPI, Tubingen - link to recording (1 hr)
Ksenia Krasileva - Evolution in Overdrive: Decoding and Encoding Genomes for Disease Resistance - Saclay Plant Science Hub invited seminar Oct 2024 - link to recording (40 min)
Chandler Sutherland - "Contributions of Mutation and Selection to the Rapid Evolution of Plant Innate Immune Receptors" - Biology of Genomes 2024 - link to recording (15 min talk plus Q&A)
Pierre Joubert - "Catching up to Fungal Plant Pathogens" - Pierre Joubert Dissertation Finishing Talk, May 15, 2023 - link to recording (1 hr seminar)
Kyungyong Seong - "Elucidating effector evolution through predicted structures" MPMI Virtual Seminar - Best MPMI Graduate Student Paper Award - link to recording (30 min seminar plus Q&A)
Ksenia - "Comparative structural genomics of pathogen effectors and plant immune receptors: from natural diversity to precise modification" UC Riverside, Sept 30th, invited seminar - link to recording (1 hr seminar)
Pierre Joubert - "Characterizing the extrachromosomal circular DNAs of Magnaporthe oryzae and their potential role in adaptation" - The 31st Fungal Genetics Conference, March 15-20 2022 - link to recording (15 min talk)
Erin Baggs - "Lessons from loss; plant immune signaling pathways and pathogen interactions from land to water" - PhD finishing talk, January 13th - link to recording (46 min seminar with intro by Ksenia)
Outreach
Chandler Sutherland - "How Plants Evolve Their Immune System" The Graduates radio podcast, hosted on on KALX 90.7 link to recording
Chandler Sutherland - "It's Not Easy Staying Green: Understanding Plant Immune Systems" Ask a Science Envoy, hosted by Wonderfest: Bay Area Beacon of Science. " link to recording
Fraces (Grace) Stark - "Fungal Immune Systems with Grace Stark" - Central Texas Mycological Society, Nov 18th 2021 7-9PM CT invited talk. Open to public with livestream on Youtube - link to recording (~1 hr 30 min with discussion)
Who we are and what we do
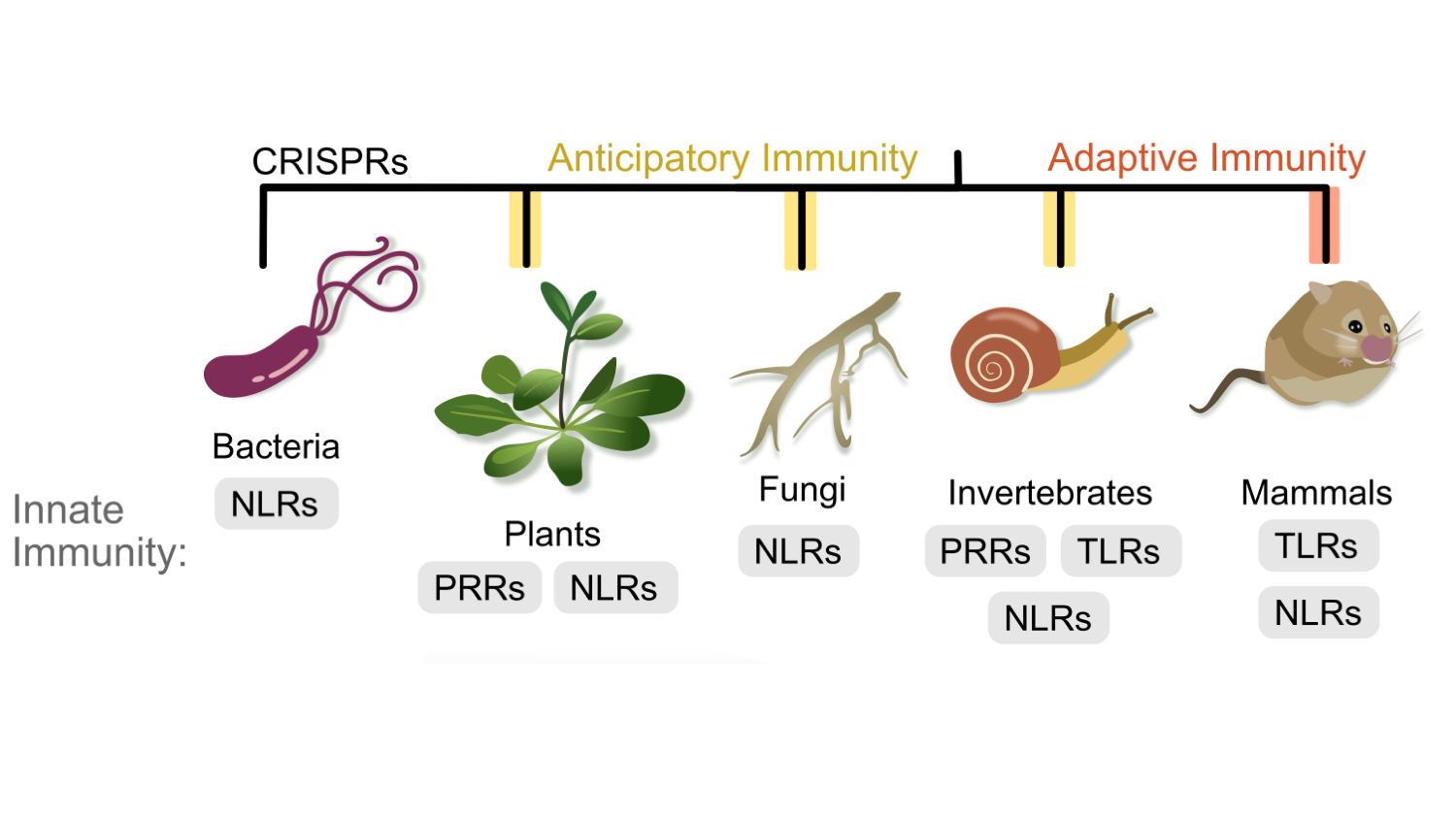
Krasileva lab an inter-disciplinary group of people. Our common goal is to understand innate immunity; currently we are working with plants and fungi. We focus on evolution and function of immune genes, as well as the mechanisms that regulate genetic diversity. To read more about specific research projects visit About us and Publications.
The lab is located in Koshland Hall, we are part of Department of Plant and Microbial Biology as well as the Innovative Genomics Institute and The Center for Computational Biology at the University of California Berkeley.
If you are interested in joining the lab:
Undergraduate research: reach out to us directly before the semester begins or look for projects at URAP and SPUR.
Graduate students: we are part of graduate programs in Microbiology and Plant Biology as well as Center for Computational Biology. To discuss a rotation, please reach out a few weeks in advance.
Postdocs: please send Ksenia your CV and a list of postdoc fellowships you would like to apply for.
Staff: occassionally, positions become available. Please, watch this space or send us a note.
Our lab culture, summarized in the Krasileva Lab Charter, is built on honesty and transparency and open communication. We aim to maintain the highest quality of research and strongest work ethic without sacrificing our health or wellbeing of our families. If you have any questions about Krasileva Lab, do not hesitate to reach out to any lab members.